Fly-QMA¶
Fly-QMA is part of the NU FlyEye platform for quantitative analysis of Drosophila imaginal discs. The package enables Quantitative Mosaic Analysis (QMA) - that is, it helps users quantify and analyze expression patterns in mosaic tissues.
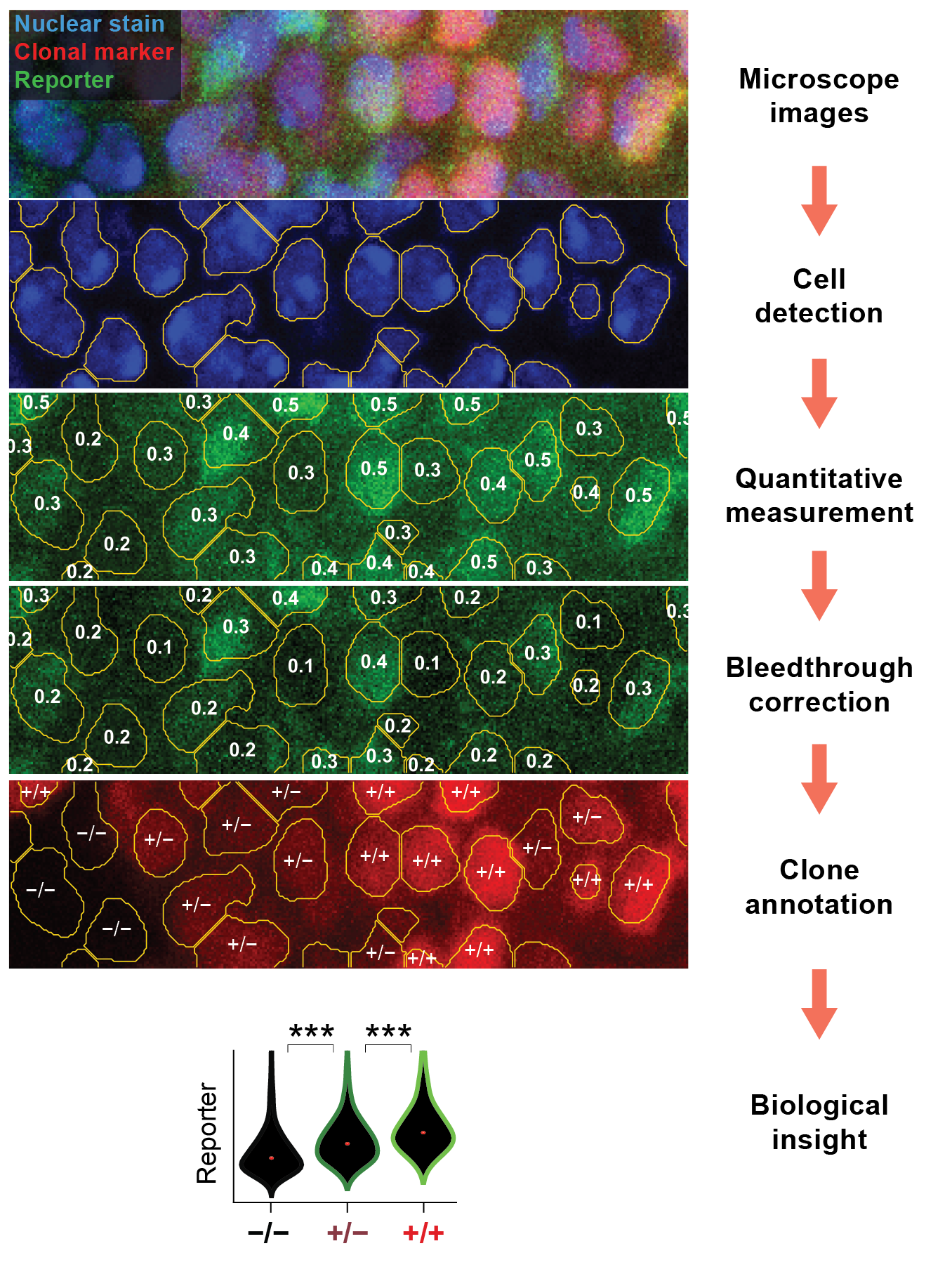
Expression patterns are typically identified by comparing the intensities of fluorescent reporters between groups of cells. Fly-QMA uses computer vision to quantify these differences in reporter expression by inferring them from microscope images. The measurements may then used to detect and analyze spatial patterns that might otherwise go unnoticed. Check out our manuscript for a detailed explanation of the algorithms underlying Fly-QMA.
Given microscopy data, Fly-QMA facilitates:
Image segmentation to detect cell nuclei
Quantitative measurement of reporter expression levels
Automated bleedthrough control for enhanced measurement accuracy
Automated annotation of clonal patch patterns
Statistical analysis of expression levels and tissue morphology
To get started, simply pip install flyqma then skim through our beginners guide.