My primary field is quantitative biology; I synthesize concepts from chemical engineering and computer science to study the behavior of biological systems. As a graduate student, I was particularly focused on exploring the chemical mechanisms that guide cells to adopt the appropriate roles and functions during the formation of complex organs.
Despite the subject matter, my role is akin to that of a modern data scientist. I combine computer vision, mathematical modeling, statistics, and dynamical systems theory to develop new analytical methods and derive meaning from experimental data. Common themes include:
- Development & validation of toy models
- Modeling stochastic processes
- Process dynamics & control
- Quantitative image analysis
- Statistical modeling & analysis
Check out the sections below for an introduction to some of my past projects, and please don’t hesitate to contact me with any ideas, suggestions, or questions!
Redundancy in gene regulatory networks
During the development of complex organisms, gene regulatory networks (GRNs) integrate external signals to ensure the timely execution of downstream events. These networks exhibit extensive functional redundancy, enabling cells to partially compensate for deleterious mutations. I sought to elucidate evolutionary forces shaping these topologies by identifying additional benefits that redundant regulatory mechanisms confer upon heterogeneous populations of organisms. My collaborators and I coupled model-based predictions with experimental validation in model organisms in an effort to generate and test new hypotheses. We showed that redundant regulatory mechanisms enable faster rates of growth and development by safeguarding against excessive protein expression when metabolic rates are high.

The results implicate a novel driving force in the evolution of genetic circuits. By adding more brakes, organisms can safely upgrade their metabolic engine in order to gain an edge in the race from embryo to adulthood.

Automated analysis of mosaic tissues
Biologists use mosaic tissues to compare the behavior of genetically distinct cells within an otherwise equivalent context. The ensuing analysis is often limited to qualitative insight. However, it is becoming clear that quantitative models are needed to unravel the complexities of many biological systems. I developed a computational framework that automates the quantification of mosaic analysis for Drosophila imaginal discs, a common setting for studies of developmental processes. The software extracts quantitative measurements from confocal images of mosaic tissues, rectifies any cross-talk between fluorescent reporters, and identifies clonally-related subpopulations of cells. Together, these functions allow users to rigorously ascribe changes in gene expression to the presence or absence of particular genes.

Cellular decisions in the developing eye
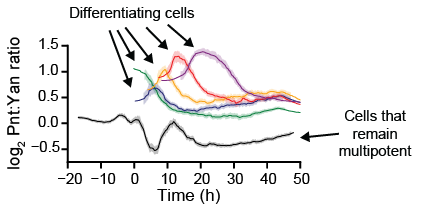
Transcription factors coordinate the timing and execution of cell differentiation by tuning the expression of target genes. Some transcription factors promote differentiation, while others impede it. Competing transcription factors are often co-expressed in vivo, and it remains unclear how cells reliably integrate their antagonistic inputs. I developed and deployed computer vision techniques to infer the expression dynamics of competing transcription factors from confocal microscope images of the developing fruit fly eye. Combined with an experimental pipeline developed by my experimental collaborators, my approach facilitates quantitative modeling and characterization of transcription factor activity both before, during, and after differentiation.